Once the list of files to upload is populated, click the green Upload All button to load your networks.ĭepending on the size and number of files queued for loading, the process might take more or less time. 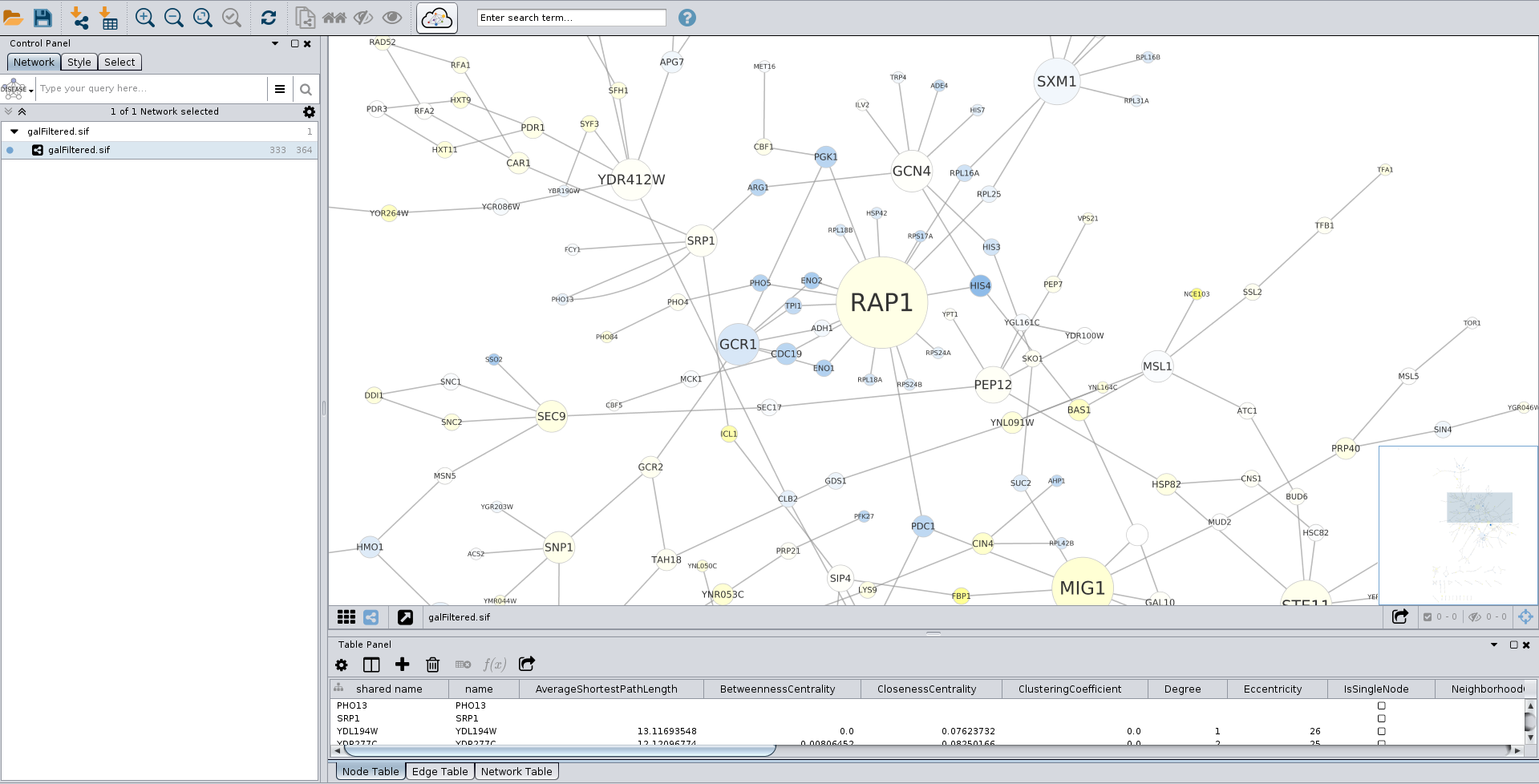
Select the desired CX file(s) from your computer.click the Upload Networks button available in your "My Account" page.To upload one or more CX networks to NDEX: Support for the upload of tabular files and other standard network formats will be added gradually in 2019. Uploading via the NDEx Web User InterfaceĪt this time, the NDEx web UI only allows the upload of networks in CX format. Please refer to the the Developer's README page for more information and links to the relevant GitHub repositories and documentation. In addition, web applications can communicate with NDEx via Javascript. Uploading ProgrammaticallyĬomputational savvy researchers and bioinformaticians can upload networks via their custom scripts using the NDEx REST API and one of the available NDEx client libraries: Java, Python or R. If you have your network in Cytoscape, it's just a couple of clicks to upload it to NDEx.Ī quick guide with screenshots is available in the CyNDEx-2 App Store page alternatively, instructions can also be found in the Cytoscape Online Manual. The latest Cytoscape 3.7 has built-in full integration with NDEx thanks to the CyNDEx-2 Core App, loading and retrieving networks form NDEx it's easy and fast. The easiest option to load your favorite networks to NDEx is to use Cytoscape. You can visit THIS PAGE for details about creating an NDEx account. Please note that a free NDEx account is required to upload and share networks in NDEx. Users can also request access to networks they do not own but they are interested in. Besides setting the network visibility, users can also decide to grant access to their network to specific NDEx users for collaborative purposes. In NDEx, users have full control on who can and cannot access their networks. There are several ways to upload and share networks in NDEx and this page provides a short review of the available options. Several case studies of Cytoscape plug-ins are surveyed, including a search for interaction pathways correlating with changes in gene expression, a study of protein complexes involved in cellular recovery to DNA damage, inference of a combined physical/functional interaction network for Halobacterium, and an interface to detailed stochastic/kinetic gene regulatory models.Last updated: December 12th, 2018 Overview The Core is extensible through a straightforward plug-in architecture, allowing rapid development of additional computational analyses and features. Cytoscape's software Core provides basic functionality to layout and query the network to visually integrate the network with expression profiles, phenotypes, and other molecular states and to link the network to databases of functional annotations. Although applicable to any system of molecular components and interactions, Cytoscape is most powerful when used in conjunction with large databases of protein-protein, protein-DNA, and genetic interactions that are increasingly available for humans and model organisms. Cytoscape is an open source software project for integrating biomolecular interaction networks with high-throughput expression data and other molecular states into a unified conceptual framework. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
